genBSDF in Windows 10
Hello community
I'm trying to use genBSDF in MS Windows 10 but I always get the same error. I've tried different computers, legacy versions of Radiance, the .exe version from Jaloxa, and the Strawberry Perl and ActivestatePerl compiler. Is this a general windows 10 issue or is it specific to me?
(I'm just using the basic blind exercise from the genBSDF tutorial. I've also tried other examples though).
The error message is below:
"
C:\Users\EVER7798\Documents\gen>genBSDF.pl +f +b -c 500 -geom inch blinds.rad 1>test_results.xml Use of uninitialized value $tmploc in scalar chomp at C:\radiance\bin\genBSDF.pl line 33. Use of uninitialized value $tmploc in concatenation (.) or string at C:\radiance\bin\genBSDF.pl line 34. Can't spawn "cmd.exe": No such file or directory at C:\radiance\bin\genBSDF.pl line 171. Could not load Radiance input
"
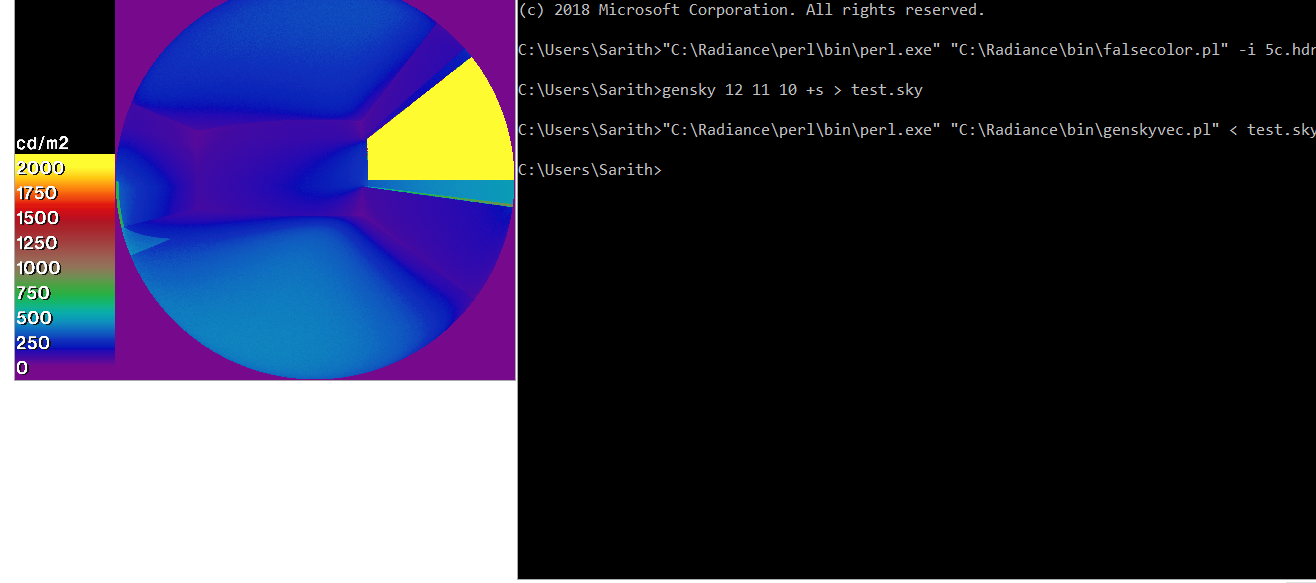
I also notice falsecolor.pl and genskyvec.pl does not work with windows 10. Is there a general issue with Perl scripts in Windows?
Thanks for any input
Update 19/06/2018
Thanks for replying sarith and rpg777, I tried what you've said and I've found that if I DO NOT assign the system path and raypath, the perl files run ok with the exception of genBSDF. (if I assign the system and ray path, only objview works).
With the paths assigned genBSDF immediately crashes. Without the paths assigns genBSDF runs a while then returns an error message.
See images below with / without the below statements:
"SET RAYPATH= .;c:\Radiance\lib" and "PATH= c:\Radiance\bin; $PATH"
I also include the text from the batch file I'm using.
Could another person try this on their windows 10 (change the file paths of course!). Has anyone else any ideas what is going wrong?
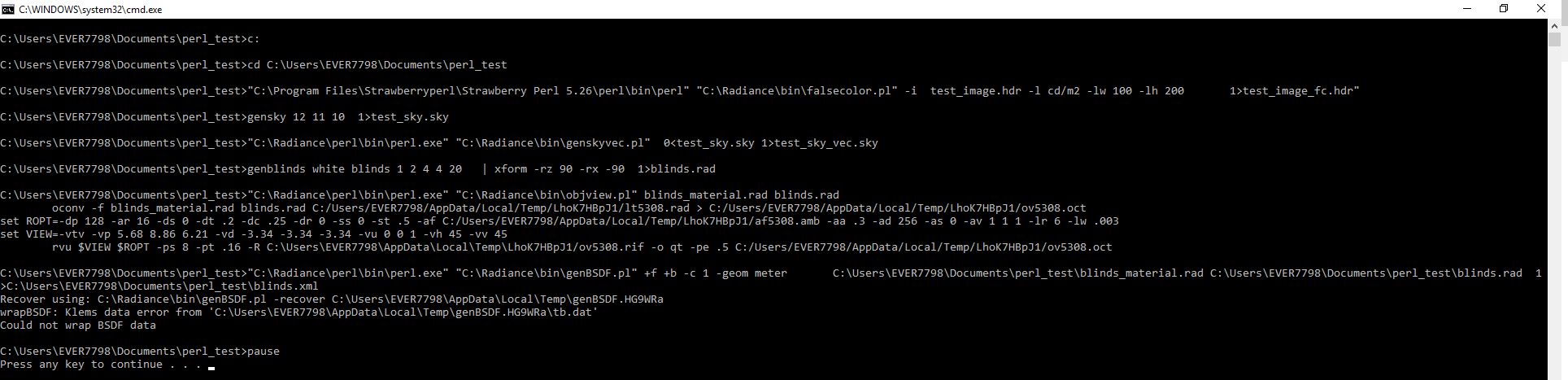
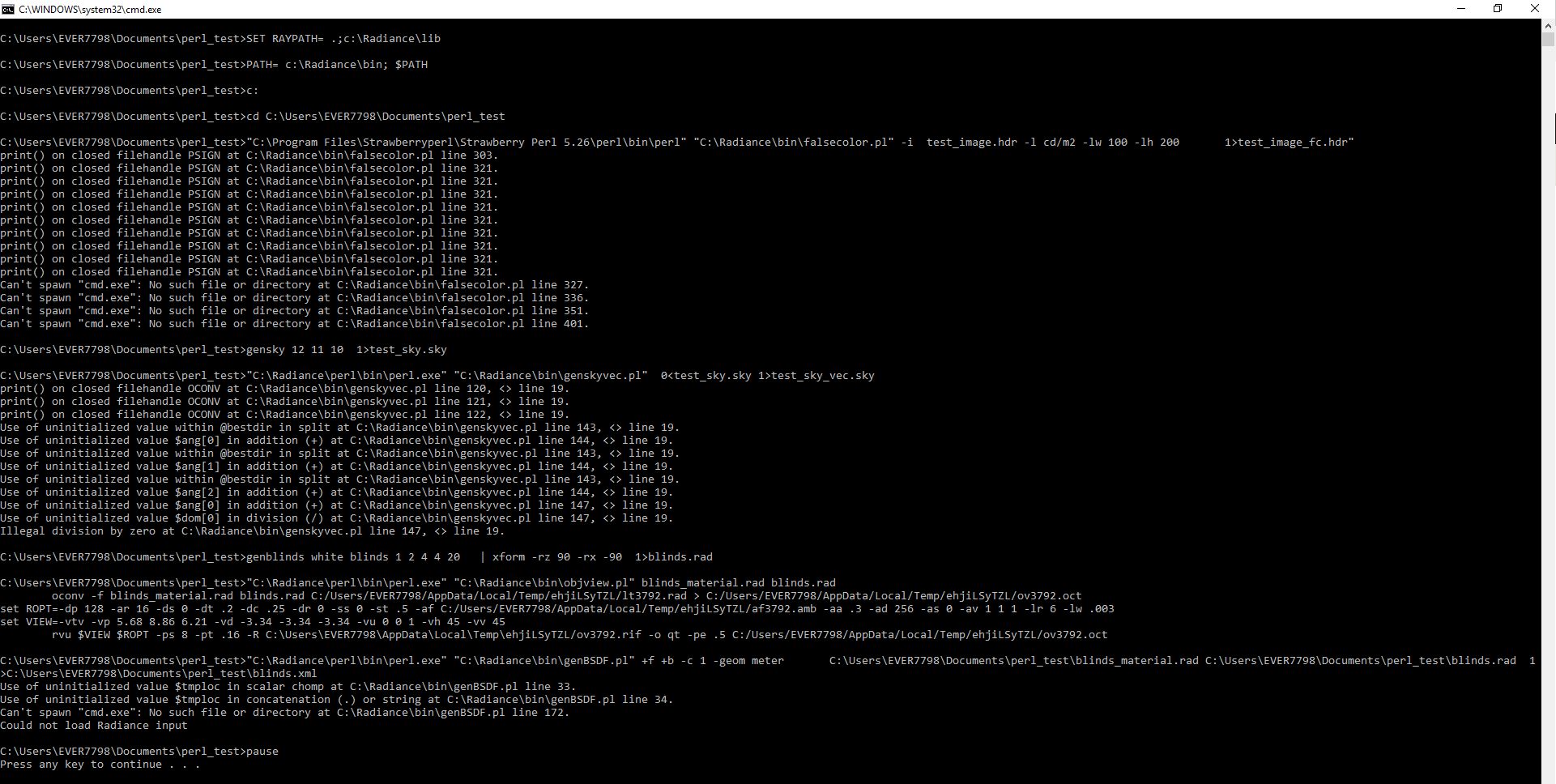
this is the batch file I'm using...
:start of batch-----------------------------------------
SET RAYPATH= .;c:\Radiance\lib PATH= c:\Radiance\bin; $PATH
c: cd C:\Users\EVER7798\Documents\perl_test
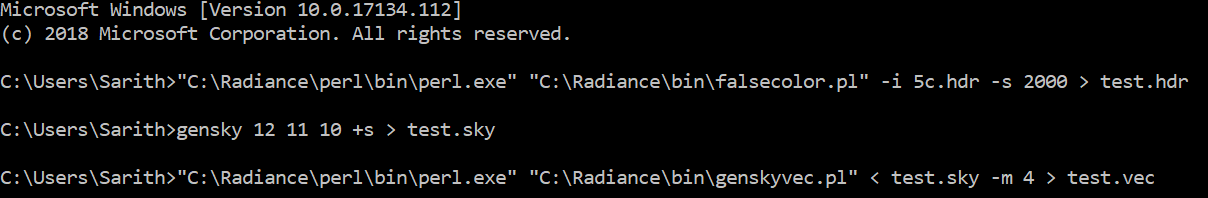
"C:\Program Files\Strawberryperl\Strawberry Perl 5.26\perl\bin\perl" "C:\Radiance\bin\falsecolor.pl" -i test_image.hdr -l cd/m2 -lw 100 -lh 200 > test_image_fc.hdr"
::---------------- gensky 12 11 10 > test_sky.sky "C:\Radiance\perl\bin\perl.exe" "C:\Radiance\bin\genskyvec.pl" < test_sky.sky > test_sky_vec.sky ::----------------
::---------------- @echo void plastic white 0 0 5 0.7 0.7 0.7 0 0 > blinds_material.rad genblinds white blinds 1 2 4 4 20 | xform -rz 90 -rx -90 > blinds.rad ::----------------
::---------------- "C:\Radiance\perl\bin\perl.exe" "C:\Radiance\bin\objview.pl" blinds_material.rad blinds.rad ::----------------
::getbbox blinds.rad
::---------------- "C:\Radiance\perl\bin\perl.exe" "C:\Radiance\bin\genBSDF.pl" +f +b -c 1 -geom meter C:\Users\EVER7798\Documents\perl_test\blinds_material.rad C:\Users\EVER7798\Documents\perl_test\blinds.rad > C:\Users\EVER7798\Documents\perl_test\blinds.xml ::----------------
pause :::end of batch-----------------------------------------------------------